Spark on EMR
High Level Problem
Our Dynamic Ads product is designed to show a user ads with specific, recommended products based on their browsing history. Each advertiser gives us a list of all of its products beforehand so that we can generate a “product catalog” for the advertiser. Sometimes, however, the product catalog that the advertiser has given us may not accurately reflect all of the products that exist on their website, and our recommendations become less accurate. Thus, we wanted to run a daily data processing job that would look at all products viewed on a given day and determine if we had a matching record in the advertiser’s product catalog. Then, we could aggregate that data to calculate a “match rate” for each advertiser, allowing us to easily target and troubleshoot troubled product catalogs.
In order to accomplish this task, we needed to perform the following actions:
- Query AdRoll’s raw logs via Presto for initial data set (8 million rows).
- For each row in the data set, query DynamoDB.
- Add each new row to a table in a PostgreSQL database (Amazon RDS).
- Summarize (reduce) the data set to get an aggregated view.
- Add each row to another aggregated table in the PostgreSQL database.
Why PySpark?
Given that we were faced with a large-scale data processing task, it became clear that we would need to use a cluster computing solution. As a new engineer who is most familiar with Python, I was initially drawn to Spark (as opposed to Hadoop) because I would be able to use PySpark, a Python wrapper around Spark. While there are other Python libraries to perform mapreduce tasks (like mrjob), PySpark was the most attractive of the Python options due to its intuitive, concise syntax and its easy maintainability. Moreover, the performance gains from using Spark were also a major contributing factor.
While the PySpark code needed to build the necessary RDDs and write to the database ended up being rather trivial, I ran into numerous operational roadblocks around deploying and running PySpark on EMR. Many of these issues stemmed from the fact that despite the new EMR 4.1.0 AMI supporting Spark and PySpark out of the box, it was difficult to find documentation on configuring Spark to optimize resource allocation and deploying batch applications. Among the various issues I encountered and ultimately solved were: installing Python dependencies, setting Python 2.7 as the default, configuring executor and driver memory, and running the spark-submit step. In this post, I present a guide based on my experience that I hope serves to smooth the development process for future users.
Spark
- Transformations vs. Actions
- Spark RDDs support two types of operations: transformations and actions. As the documentation clearly explains, transformations (like
map) create a new dataset from an existing one, but they are lazy, meaning that they do not compute their results right away. In other words, transformations simply remember which transformations were applied to the base data. Actions (likeforeachPartition), on the other hand, return a value to the driver after running a computation. The computations associated with a transformation are not actually computed until an action is called. - An understanding of this difference is hugely important in actually leveraging Spark’s performance gains. For example, prior to understanding this crucial difference, I was actually making the 8 million DynamoDB calls twice, instead of just once. With this understanding, I was able to decrease my job runtime by more than 50%.
- Below, I have shown the difference between the code before and after this realization. Depending on the size of your data set and the memory available, you can either call
.persist()on the RDD that you will reuse (rdd1in our case) or write the data to disk. Due to the size of our data set and the fact that the intermediary RDD was precisely what I wanted to store in the database, I chose to writerdd1to the database, and then query the database to initiate a new RDD.
- Spark RDDs support two types of operations: transformations and actions. As the documentation clearly explains, transformations (like
##### INEFFICIENT CODE #####
base_rdd = sc.parallelize(presto_data)
# We do not actually query DynamoDB until foreachPartition is called,
# as map is a transformation and foreachPartition is an action.
rdd1 = base_rdd.map(query_dynamo_db)
rdd1.foreachPartition(add_to_db)
# At this point, because query_dynamo_db was within a map function
# (a transformation, which is remembered by rdd1), we actually query
# DynamoDB all over again when foreachPartition is called.
rdd2 = rdd1.map(lambda x: (x[0], x[1])).reduceByKey(lambda x,y: x+y)
rdd2.foreachPartition(add_to_db)
##### IMPROVED CODE #####
base_rdd = sc.parallelize(presto_data)
rdd1 = base_rdd.map(query_dynamo_db)
rdd1.foreachPartition(add_to_db)
# Here we build the second RDD based on the data which was added
# to the database in the first foreachPartition action, instead of
# based upon rdd1. Thus, we do not repeat the DynamoDB calls.
db_rows = get_rows_from_db()
new_rdd = sc.parallelize(db_rows)
rdd2 = new_rdd.map(lambda x: (x[0], x[1])).reduceByKey(lambda x,y: x+y)
rdd2.foreachPartition(add_to_db)
- Partitions
- An important parameter when considering how to parallelize your data is the number of partitions to use to split up your dataset.
- Spark runs one task (action) per partition, so it’s important to partition your data appropriately.
- Spark’s documentation recommends 2-4 partitions for each CPU in the cluster.
- RDDs created from
textFileorhadoopFilewill use the mapreduce API to determine the number of partitions and thus will have a reasonable predetermined number of partitions. - On the other hand, the data set returned from my Presto query returns a number of partitions determined by the
spark.default.parallelismsetting. By default, this setting sets the number of partitions to the number of executors (spark.executor.instances, 20 in our case) or 2, whichever is greater. Thus, in order to optimize, we manually tuned the number of partitions to be equal to 80 (20 executors * 4 partitions each).
base_rdd = sc.parallelize(presto_data, 80)
- Importing from Self-created Modules
- Unlike in base Python, where one is able to import from any other Python file within a given repository, in Pyspark, external self-created libraries need to be zipped up and then added to the global SparkContext object.
- First the SparkContext object must be created, and then the zip file is added to the object via the addPyFiles method.
from pyspark import SparkContext
sc = SparkContext()
sc.addPyFile('/home/hadoop/spark_app.zip')
- Using Sqlalchemy
- Because of the distributed nature of Spark, a Sqlalchemy
Sessionobject can not be shared across multiple executors or paritions. - For this reason, a new
Sessionobject must be created per partition, andsqlalchemy.create_engineandsqlalchemy.orm.sessionmakermust be imported within each partion. All of this is done within a function called byforeachPartition.
- Because of the distributed nature of Spark, a Sqlalchemy
def add_to_db(iterator):
from sqlalchemy import create_engine
from sqlalchemy.orm import sessionmaker
engine = create_engine(DATABASE_URL)
session = sessionmaker(bind=engine)
Session = session()
for obj in iterator:
Session.add(obj)
Session.commit()
my_rdd.foreachPartition(add_to_db)
EMR
- Packaging & Deploying Code
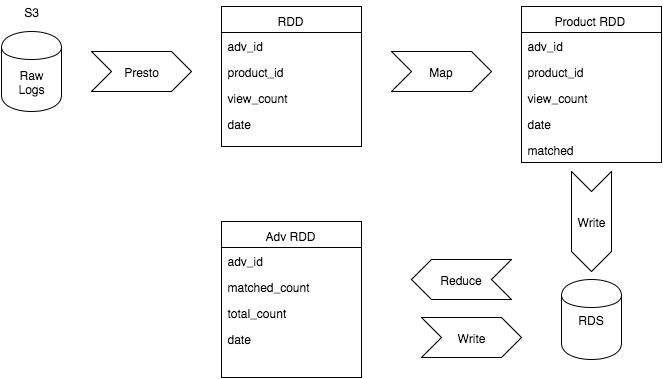
- The PySpark code was organized as follows:
- Use Presto to query raw logs on S3 each day for the count (
view_count) of product views per product_id. - Convert the results of this query into an RDD.
- Using Spark, map over this RDD to query DynamoDB to see if each row in the RDD has a matching record in DynamoDB. The result of this mapping function is a new RDD (named “Product RDD” in the diagram below), which now has a boolean field
matchedto indicate whether or not a matching record was found. - Write the contents of this new RDD to the Postgres database, an AWS RDS instance.
- Reduce this data to the advertiser level, giving a summary of the number of matched views vs. total views by advertiser by day.
- Use Presto to query raw logs on S3 each day for the count (
- As discussed above, in order to access self-created Python modules from within my main PySpark script, the modules had to be zipped up. My solution to this was to create a main module called
spark_app, within which I had sub-modulesmodelandutils. I then created adeploy.pyscript that serves to zip up thespark_appdirectory and push the zipped file to a bucket on S3. - Similarly, the
deploy.pyscript also pushesprocess_data.py(the main PySpark script) andsetup.sh(the bootstrap action script) to the same bucket on S3. - Bootstrap actions are run before your steps run. In this way, they are used to set up your cluster appropriately. For our purposes, that involved installing my necessary Python dependencies and copying
spark_app.zipandprocess_data.pyto the right directory. - Below is our
setup.shscript:
- The PySpark code was organized as follows:
#!/bin/bash
# Install our dependencies - replace these libraries with your needs!
sudo yum install -y gcc python-setuptools python-devel postgresql-devel
sudo python2.7 -m pip install SQLAlchemy==1.0.8
sudo python2.7 -m pip install boto==2.38.0
sudo python2.7 -m pip install funcsigs==0.4
sudo python2.7 -m pip install pbr==1.8.1
sudo python2.7 -m pip install psycopg2==2.6.1
sudo python2.7 -m pip install six==1.10.0
sudo python2.7 -m pip install wsgiref==0.1.2
sudo python2.7 -m pip install requests[security]
# Download code from S3 and set up cluster
aws s3 cp s3://your-bucket/spark_app.zip /home/hadoop/spark_app.zip
aws s3 cp s3://your-bucket/process_data.py /home/hadoop/process_data.py
- Running the Job
- Ultimately, I chose to run this Spark application using a scheduled Lambda function on AWS (discussed below), but for the purposes of this blog post, I have included both the Lambda function which utilizes boto3 and the equivalent cluster launch using the
aws cli. - AWS Lambda Function:
- Ultimately, I chose to run this Spark application using a scheduled Lambda function on AWS (discussed below), but for the purposes of this blog post, I have included both the Lambda function which utilizes boto3 and the equivalent cluster launch using the
import boto3
def lambda_handler(json_input, context):
client = boto3.client('emr', region_name='us-west-2')
client.run_job_flow(
Name='YourApp',
ReleaseLabel='emr-4.1.0',
Instances={
'MasterInstanceType': 'm3.xlarge',
'SlaveInstanceType': 'm3.xlarge',
'InstanceCount': 21,
'Ec2KeyName': 'ops',
'KeepJobFlowAliveWhenNoSteps': False,
'TerminationProtected': False,
'Ec2SubnetId': 'subnet-XXXX
},
Steps=[
{
'Name': 'YourStep',
'ActionOnFailure': 'TERMINATE_CLUSTER',
'HadoopJarStep': {
'Jar': 'command-runner.jar',
'Args': [
'spark-submit',
'--driver-memory','10G',
'--executor-memory','4G',
'--executor-cores','4',
'--num-executors','20',
'/home/hadoop/process_data.py'
]
}
},
],
BootstrapActions=[
{
'Name': 'cluster_setup',
'ScriptBootstrapAction': {
'Path': 's3://your-bucket/subfolder/setup.sh',
'Args': []
}
}
],
Applications=[
{
'Name': 'Spark'
},
],
Configurations=[
{
"Classification": "spark-env",
"Properties": {
},
"Configurations": [
{
"Classification": "export",
"Properties": {
"PYSPARK_PYTHON": "/usr/bin/python2.7",
"PYSPARK_DRIVER_PYTHON": "/usr/bin/python2.7"
},
"Configurations": [
]
}
]
},
{
"Classification": "spark-defaults",
"Properties": {
"spark.akka.frameSize": "2047"
}
}
],
VisibleToAllUsers=True,
JobFlowRole='EMR_EC2_DefaultRole',
ServiceRole='EMR_DefaultRole'
)
- AWS CLI Equivalent:
aws emr create-cluster --name "YourApp" --release-label emr-4.1.0 \
--use-default-roles \
--ec2-attributes KeyName=YourKey,SubnetId=YourSubnet \
--applications Name=Spark \
--configurations https://bucket.s3.amazonaws.com/key/pyspark_py27.json \
--region us-west-2 --instance-count 21 --instance-type m3.xlarge \
--bootstrap-action Path="s3://your-bucket/path/to/script" \
--steps Type=Spark,Name="StepName",ActionOnFailure=TERMINATE_CLUSTER,\
Args=[--driver-memory,10G,\
--executor-memory,4G,\
--executor-cores,4,\
--num-executors,20,\
/home/hadoop/process_data.py]
- Notes on Parameters:
- Configurations
- Because we were also accessing data from Presto within the main Pyspark script, it was necessary that all of our machines were using Python 2.7 as the default (Hive/Presto requirement). This involved configuring
PYSPARK_PYTHONandPYSPARK_DRIVER_PYTHONenvironment variables. - Additionally, we needed to add a special configuration to increase the
spark.akka.frameSizeparameter. The maximum frame size is 2047, so that is what we have set it to. This may be unnecessary for some applications, but if you receive an error about frameSize, you will know that you need to add in a configuration for this parameter. - Note: the
--configurationsparameter in the AWS CLI example simply provides a url to a json file stored on S3. That file should contain the json blob fromConfigurationsin the boto3 example above.
- Because we were also accessing data from Presto within the main Pyspark script, it was necessary that all of our machines were using Python 2.7 as the default (Hive/Presto requirement). This involved configuring
- Steps
- Within the Spark step, you can pass in Spark parameters to configure the job to meet your needs. In our case, I needed to increase both the driver and executor memory parameters, along with specifying the number of cores to use on each executor. We also found that we needed to explicitly stipulate that Spark use all 20 executors we had provisioned.
- The final argument in the list is the path to the main PySpark script (
process_data.py) that runs the actual data processing job.
- Applications
- You must explicitly specify that you intend to run a Spark application.
- Configurations
- Scheduling the Job
- With the recent announcement of Amazon’s newly available ‘Scheduled Event’ feature within Lambda, we were able to very easily set up a recurring cluster launch on a cron schedule.
- As shown above, the code to launch the cluster was extremely simple, leveraging the power of boto3’s
run_job_flowmethod. - The only wrinkle here was properly configuring the role for the lambda function, so that it would have the right permissions to launch an EMR cluster. It was necessary that the action “elasticmapreduce:RunJobFlow” had the “iam:PassRole” role associated with it.
- Below is the policy for the role:
{
"Version": "2012-10-17",
"Statement": [
{
"Effect": "Allow",
"Action": [
"logs:CreateLogGroup",
"logs:CreateLogStream",
"logs:PutLogEvents"
],
"Resource": "arn:aws:logs:*:*:*"
},
{
"Effect": "Allow",
"Action": [
"elasticmapreduce:RunJobFlow",
"iam:PassRole"
],
"Resource": "*"
}
]
}
Closing Thoughts
While there were several hurdles to overcome in order to get this PySpark application running smoothly on EMR, we are now extremely happy with the successful and smooth operation of the daily job. I hope this guide has been helpful for future PySpark and EMR users.
Finally, after taking a one-day Spark course, I have realized that for performance reasons, it likely makes sense to use Spark’s dataframes instead of RDDs for some parts of this job. Stay tuned for a future post on transitioning to dataframes and (hopefully) resulting performance gains!